Research
RNA Biology & Regeneration Lab
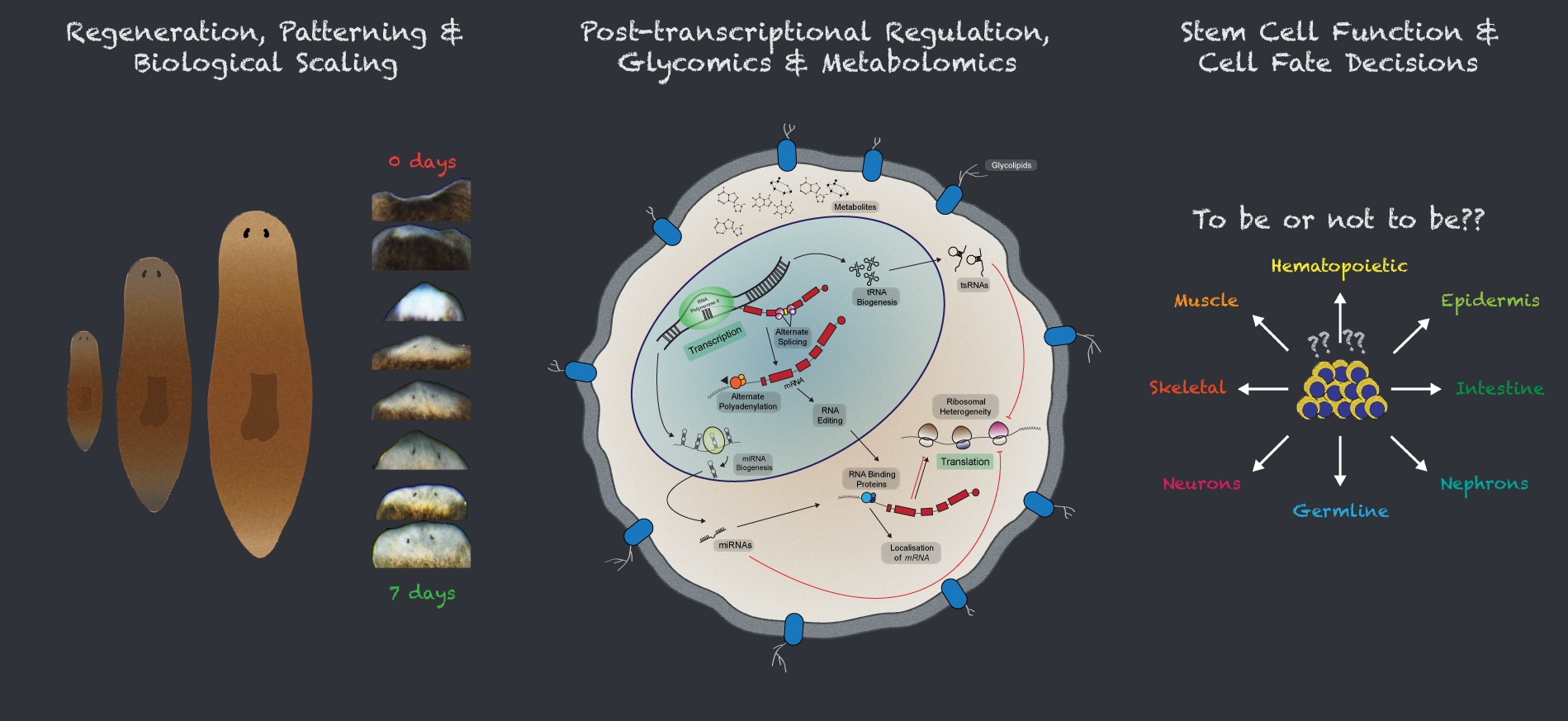
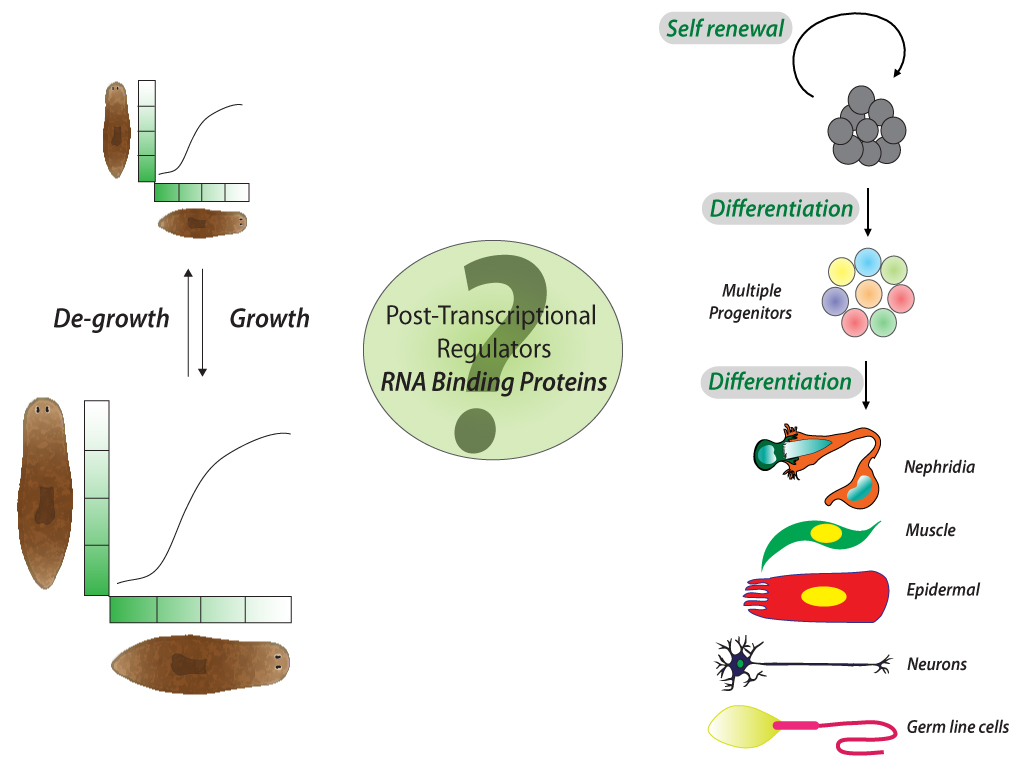
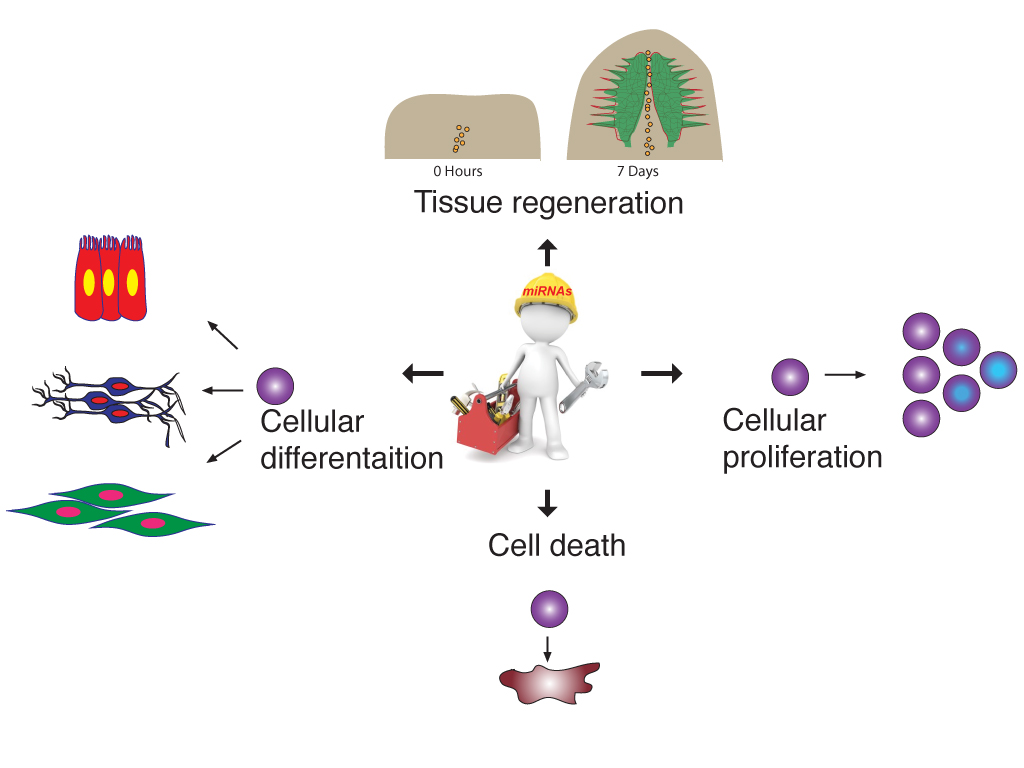
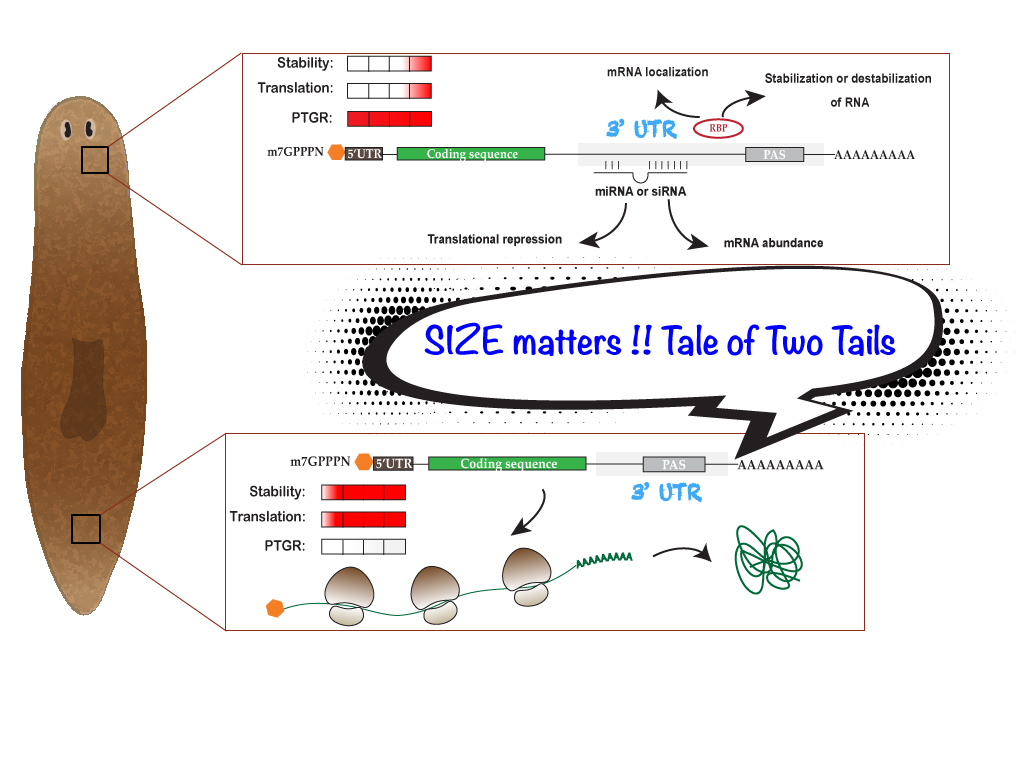
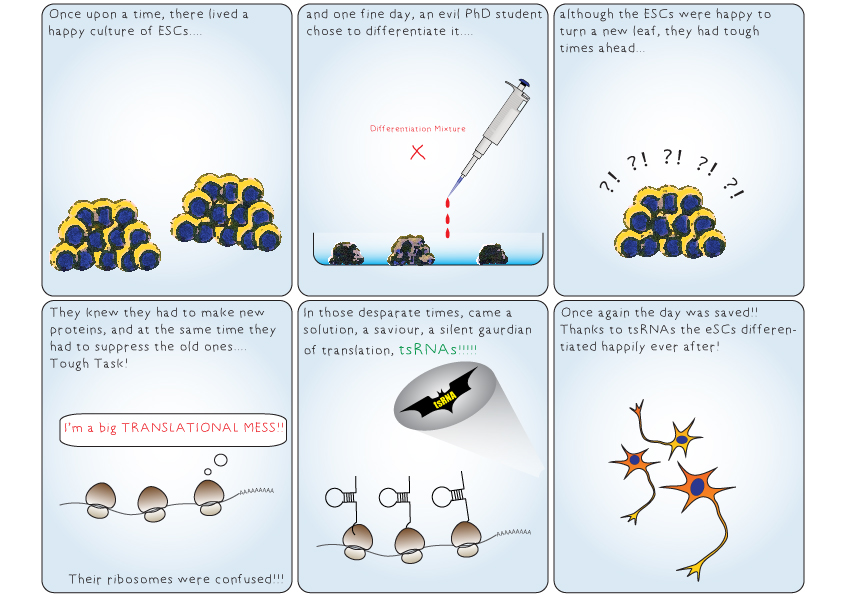
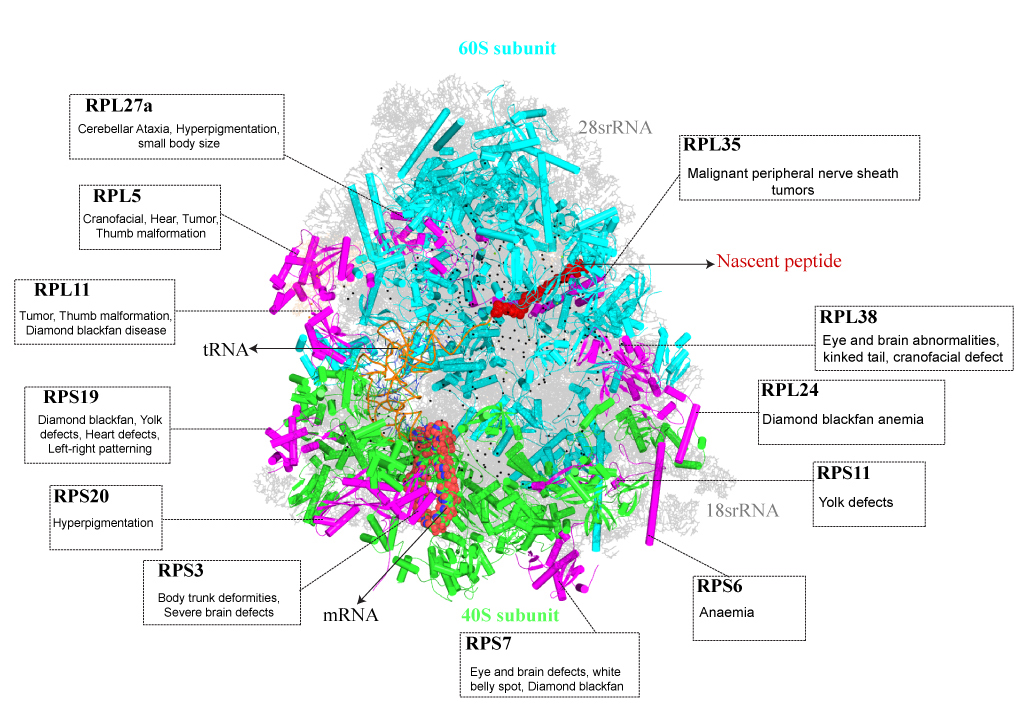
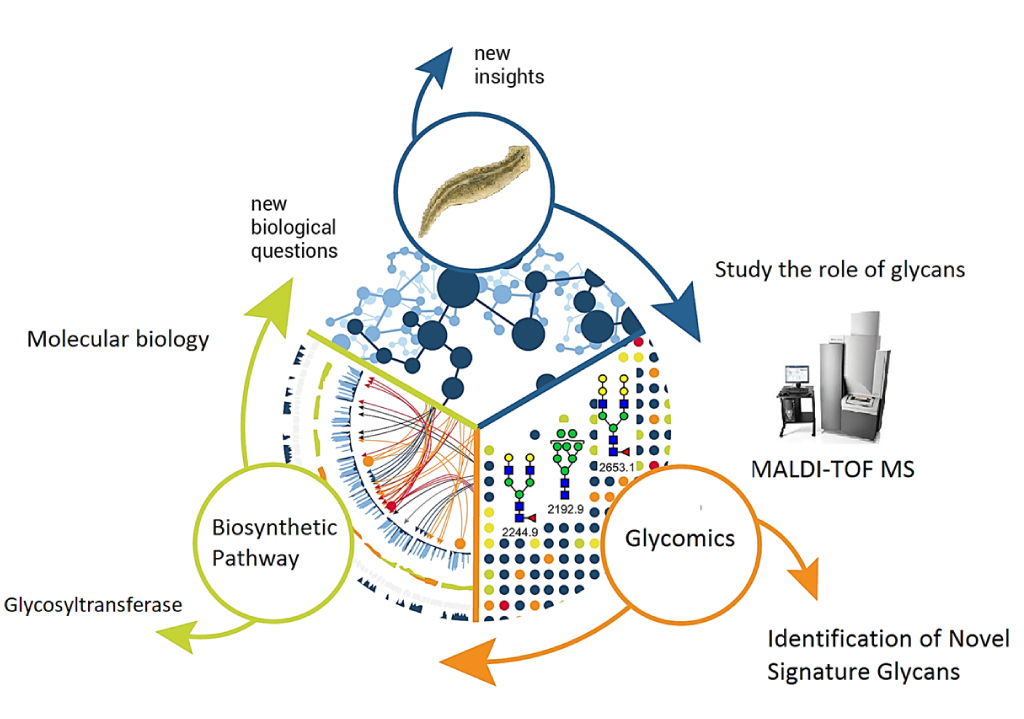
A cell undergoes dynamic changes during its transition from one cell state to another, which happens during both tissue morphogenesis and stem cell function. However, the underlying mechanisms critical for these changes remain poorly understood. Our lab is interested in exploring the regulatory networks that fine-tune gene expression, such as post-transcriptional gene regulation. Often, the mere expression of an mRNA does not relate to its functionality, which suggest presence of certain “determinants” that dictate when and where these mRNAs should be translated. These regulatory networks involve coordination between miRNAs, several RNA binding proteins and various other RNA species. We employ a whole organism (using planarian Schmidtea mediterranea regeneration) approach coupled with in vitro data (embryonic stem cell differentiation) to understand these regulatory networks. Further, we employ a metabolomics and glycomics approach to understand regeneration and cell state transitions.
About Us

Learn About our Team & Research.
Principal Investigator
Graduate Students

Nivedita Chaudhary 
niveditac at instem dot res dot in
Education: MSc Medical Biotechnology from All India Institute of Medical Sciences, New Delhi.
Our project delves into the role of Tudor proteins in stemness and regeneration, expanding their study from germline to somatic stem cells. Using planaria and mouse embryonic stem cells as models, we aim to unravel the proteins' functions in these contexts.

Atriya Mazumdar 
matriya at instem dot res dot in
Education: MSc biotechnology from RGCB, Trivandrum.
Our research focuses on understanding the role of Poly A Tail length in mRNA localization, bridging a critical gap in existing knowledge. While much is known about its influence on stability and translation, the mechanism governing localization remains unclear. Using planaria as model system, we aim to address this question.

Nikhil Kumar Jaligam 
nikhilkk at instem dot res dot in
Education: M.Sc from Osmaina University.
I am interested in the fields of regeneration and stem cell biology, exploring the translational regulation of stem cells and the pivotal role of ribosomes in this process. By unraveling the intricate mechanisms governing cellular rejuvenation, I aim to contribute to advancements in regenerative medicine and therapeutic interventions.

Sharon Samuel 
sharonsamuel at instem dot res dot in
Education: M. Pharm from Ramaiah University of Applied Sciences.
I am interested in the molecular aspects of regeneration, in Planarian flatworms, and how stem cells interact in an in vivo system.

Namita Mukundan 
namitam at instem dot res dot in
Education: MSc Biotechnology from Amrita School of Biotechnology , Kerala.
Understanding the role of PolyA Binding Protein (Nuclear) in planarian homeostasis & regeneration.
Post-Doctoral Fellows
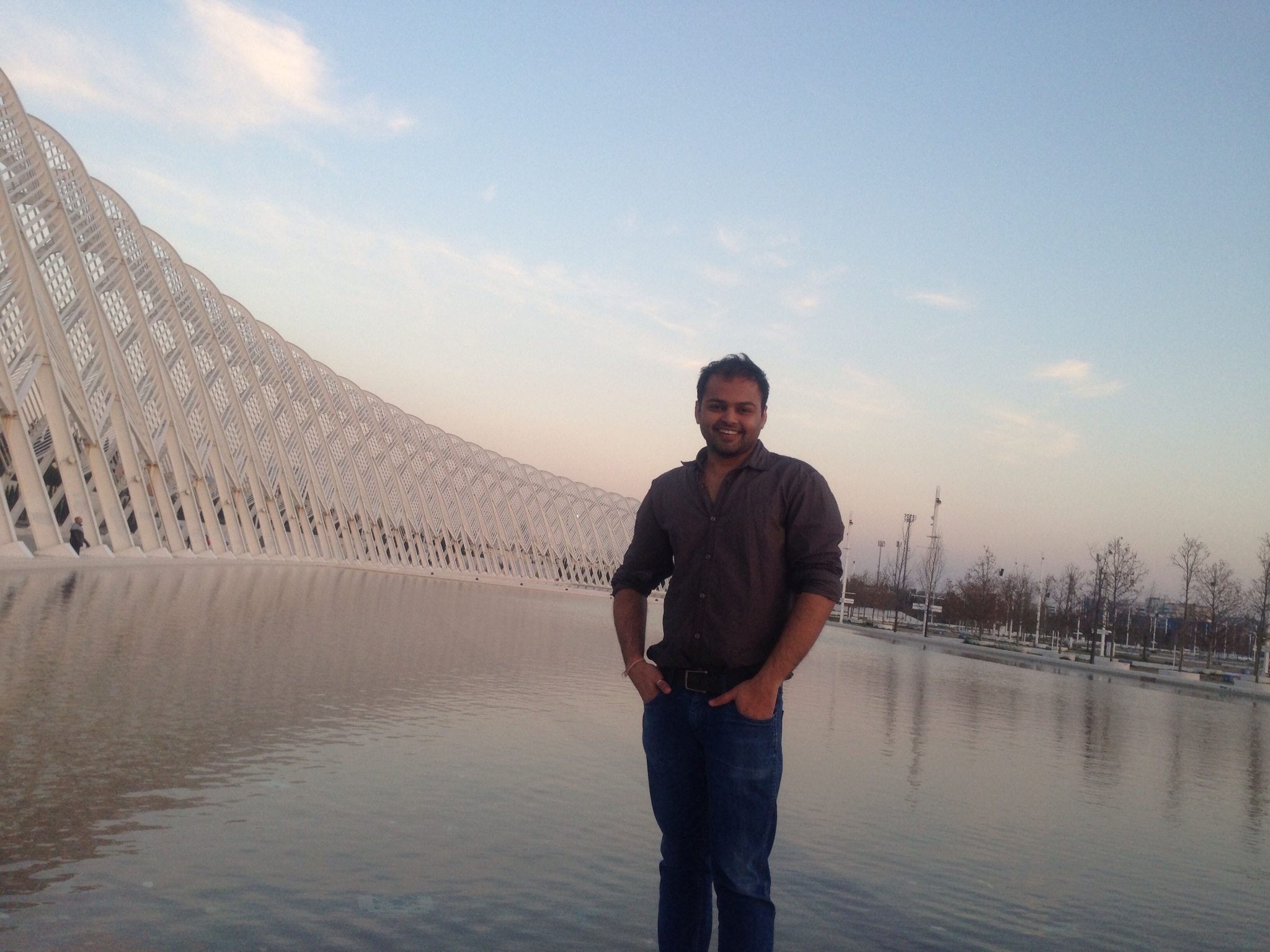

Ankit Arora 
aankit at instem dot res dot in
Education: Phd from University of Cologne, Germany.
Next-generation sequencing (NGS) data analysis
Junior Research Fellows
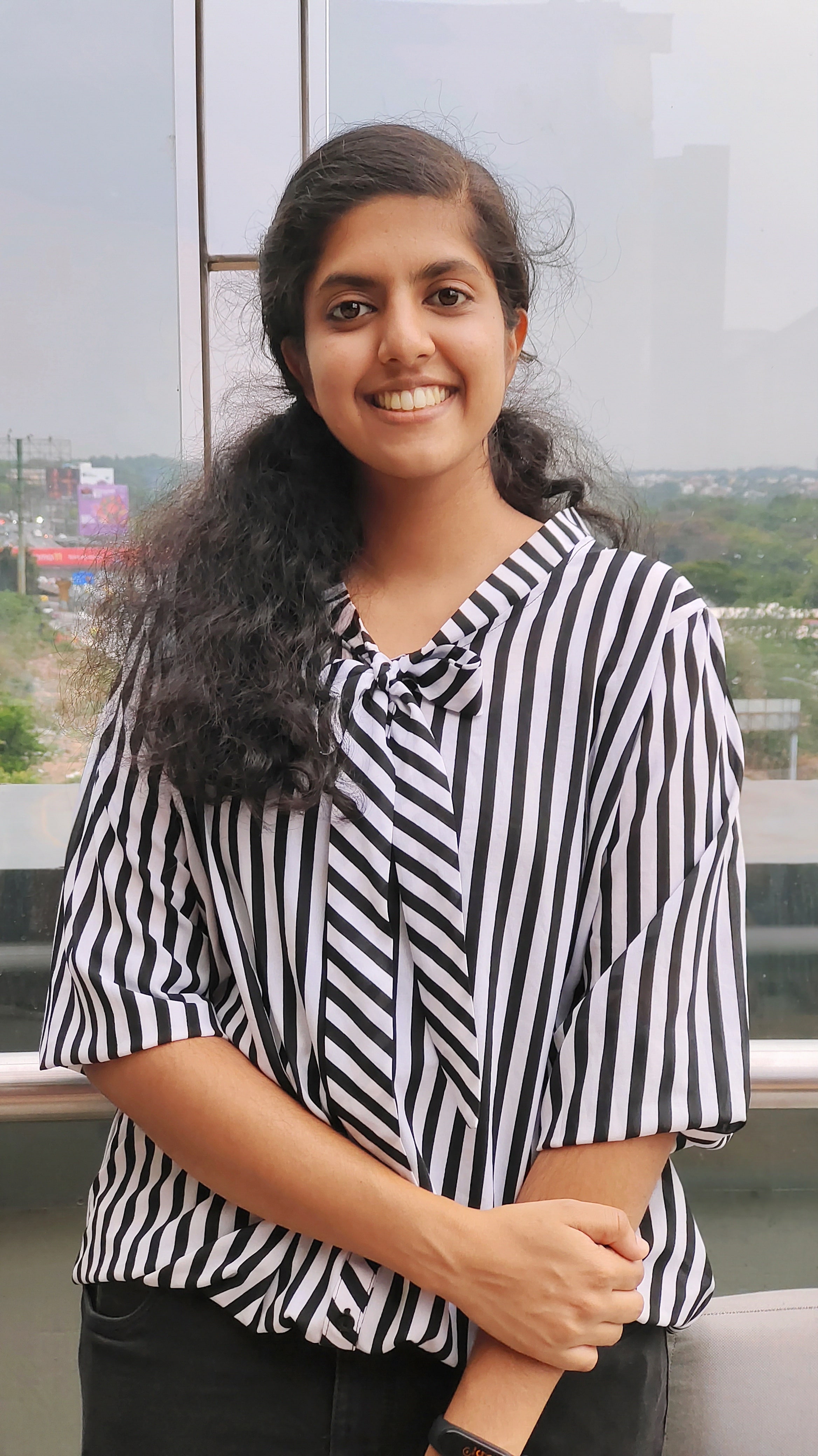

Swathi Pavithran 
swathipavithran at instem dot res dot in
Education: M.Sc Biotechnology from Christ University, Bangalore.
Our project focuses on exploring the translational control mechanisms that govern stem cell state transitions using the planarian species, Schmidtea mediterranea as a model system. Specifically, we are investigating the role of ribosomal proteins in translational regulation during cellular differentiation.
Alumni
| Niyanta Kumar | Junior Research Fellow | 2010-2012 | PhD Student University of Wisconsin-Madison |
|---|---|---|---|
| Srishti Baid | Junior Research Fellow | 2015-2017 | PhD Student University of kansas |
| Viraj Doddihal | Masters dissertation Student | 2014-2015 | PhD Student Stowers Institute for Medical Research |
| Giulia | iFOM Exchange Student | 2015-2016 | Masters Student iFOM, Italy |
| Aparna Nair | Junior Research Fellow | 2012-2014 | PhD Student CSIR Neet, Nagpur |
| Prathyusha Pavanram | Junior Research Fellow | 2012-2014 | Research Assistant Uniklinik RWTH, Aachen, Germany |
| Divya Sridhar | Junior Research Fellow | 2014-2016 | PhD Student University of Oxford |
| Pranavi Dasari | Junior Research Fellow | 2012-2014 | Agastya International Foundation Bangalore |
| Deepak Poduval Balakrishnan | Junior Research Fellow | 2012-2014 | PhD Student University of Bergen |
| Debayan Bose | Junior Research Fellow | 2012-2013 | PhD Student Academia Sinica |
| Davide Saroni | iFOM Exchange Student | 2015-2016 | Completed Masters |
| Vidyanand Sasidharan | Graduate Student | 2012-2017 | Post Doctral Fellow Sánchez Lab, Stowers Institute for Medical Research |
| Dhiru Bansal | Graduate Student | 2012-2017 | Post Doctral Fellow Miska Lab, Gurdon Institute, Cambridge |
| Praveen Anand | Post Doctoral Fellow | 2015-2016 | Post Doctrol Fellow Dana Farber Cancer Institute |
| Rosana Babu | Post Doctrol Fellow | 2016-2017 | Post Doctrol Fellow University of Hong Kong (HKU) |
| Anna Protasio | Post Doctrol Fellow | 2016-2018 | Post Doctorol Fellow University of Cambridge and Wellcome Trust Sanger Institute, UK. |
| Kaushik Iyer | Junior Research Fellow | 2017-2018 | Pandorum Technologies, CCAMP, Bangalore. |
| Arunabha Sarkar | Senior Research Fellow | 2017-2018 | |
| Nishtha Nayyar | Post Doctoral Fellow | 2016-2018 | Principal Investigator NBAIR, Bangalore. |
| Jahnavi Kulkarni | Junior Research Fellow | 2015-2019 | Biomarker Scientist BioMarkIQ |
| Rajdeep Das | Post Doctoral Fellow | 2017-2019 | Post Doctoral Fellow NCBS, Bangalore. |
| Sabarinath PS | Post Doctoral Fellow | 2014-2017 | Post Doctoral Fellow University of Nebraska Medical Center. |
| Sai Sowndarya | Junior Research Fellow | 2018-2020 | Graduate Student York University, Canada. |
| Srikar Krishna | Graduate Student | 2011-2021 | Post Doctoral Fellow Yale University. |
| Mohamed Haroon | Graduate Student | 2017-2021 | Post Doctoral Fellow inStem, Bangalore. |
| Vairavan Lakshmanan | Graduate Student | 2013-2021 | Post Doctoral Fellow NCCS-GIS, Singapore. |
| Souradeep Sarkar | Graduate Student | 2016-2022 | Post Doctoral Fellow Stanford University. |
| Venitha Bernard | Junior Research Fellow | 2021-2022 | Graduate Student University of Pennsylvania. |
| Nivitha Murali | Junior Research Fellow | 2019-2020 | Graduate Student University of Pennsylvania |
| Navjoth Menon | Junior Research Fellow | 2020-2021 | Eurofins IT delivery center. |
| Sheetal Ambardar | Post Doctoral Fellow | 2018-2019 | Assistant Professor University of Jammu |
| Vinay Kumar Dubey | Graduate Student | 2017-2024 | Post Doctoral Fellow Duke University |
| Nivedita Hariharan | Graduate Student | 2017-2024 | Setting up a Startup |
Publications
- Giulia I. Corsi, Veerendra P. Gadekar, Henriette Haukedal, Nadezhda T. Doncheva, Christian Anthon, S. Ambardar, Palakodeti D , P. Hyttel, K. Freude, S. Seemann, J. Gorodkin. The transcriptomic landscape of neurons carrying PSEN1 mutations reveals changes in extracellular matrix components and non-coding gene expression. Neurobiology of Disease December 2022. DOI: 10.1016/j.nbd.2022.105980

- Vinay Kumar Dubey, Souradeep R. Sarkar, Vairavan Lakshmanan, Rimple Dalmeida, A. Gulyani, Palakodeti D. Smed-ETS-1 regulates cathepsin+ cell function and epidermal lineage landscape via basement membrane remodeling. Journal of cell science September 2022. DOI: 10.1242/jcs.259900

- Souradeep R. Sarkar, Vinay Kumar Dubey, Anusha Jahagirdar, Vairavan Lakshmanan, M. Haroon, S. Sowndarya, R. Sowdhamini, Palakodeti D. DDX24 is required for muscle fiber organization and the suppression of wound-induced Wnt activity necessary for pole re-establishment during planarian regeneration. Developmental biology May 2022. DOI: 10.1016/j.ydbio.2022.04.011

- Nivedita Hariharan, Sumana Ghosh, Palakodeti D. The story of rRNA expansion segments: Finding functionality amidst diversity. Wiley Interdisciplinary Reviews: RNA April 2022. DOI: 10.1002/wrna.1732

- M. Haroon*, P. Vemula*, Palakodeti D. Flow Cytometry Analysis of Planarian Stem Cells Using DNA and Mitochondrial Dyes. Bio-protocol 2022. DOI: 10.21769/BioProtoc.4299

- Pratul Kumar Jain, Shashank Jayappa, T. Sairam, Anupam Mittal, Sayan Paul, Vinay J. Rao, Harshil Chittora, D. Kashyap, Palakodeti D , K. Thangaraj, J. Shenthar, Rakesh Koranchery, Ranjith Rajendran, Haghighi Alireza, K. Mohanan, A. Rathinavel, Perundurai S. Dhandapany. Ribosomal protein S6 kinase beta-1 gene variants cause hypertrophic cardiomyopathy. Biology, Medicine December 2021. DOI: 10.1136/jmedgenet-2021-107866

- Michelle Ninochka D’Souza, Sarayu Ramakrishna, B. K. Radhakrishna, Vishwaja Jhaveri, Sreenath Ravindran, Lahari Yeramala, Palakodeti D , Ravi S Muddashetty. Function of FMRP domains in regulating distinct roles of neuronal protein synthesis. Molecular Neurobiology October 2022. DOI: 10.1007/s12035-022-03049-1

- Sameena Nikhat, A. D. Yadavalli, Arpita Prusty, Priyanka Narayan, Palakodeti D , C. Murre, J. Pongubala. A regulatory network of microRNAs confers lineage commitment during early developmental trajectories of B and T lymphocytes. Proceedings of the National Academy of Science November 2021. DOI: 10.1073/pnas.2104297118

- Oindrila Bhattacharjee, Uttkarsh Ayyangar, Ambika S. Kurbet, Vairavan Lakshmanan, Palakodeti D , F. Ginhoux, Srikala Raghavan. Epithelial-Macrophage Crosstalk Initiates Sterile Inflammation in Embryonic Skin. Frontiers in Immunology October 2021. DOI: 10.3389/fimmu.2021.718005

- Anirudh Chakravarthy, Anirudh Nandakumar, Geen George, Shyamsundar Ranganathan, Suchitta Umashankar, Nishan Shettigar, Palakodeti D , A. Gulyani, A. Ramesh. Engineered RNA biosensors enable ultrasensitive SARS-CoV-2 detection in a simple color and luminescence assay. Life Science Alliance September 2021. DOI: 10.26508/lsa.202101213

- S. P. Subramanian, Vairavan Lakshmanan, Palakodeti D , R. Subramanian. Glycomic and glycotranscriptomic profiling of mucin-type O-glycans in planarian Schmidtea mediterranea.. Glycobiology Septembeer 2021. DOI:10.1093/glycob/cwab097

- Nishan Shettigar, Anirudh Chakravarthy, Suchitta Umashankar, Vairavan Lakshmanan, Palakodeti D , A. Gulyani. Discovery of a body-wide photosensory array that matures in an adult-like animal and mediates eye–brain-independent movement and arousal. Proceedings of the National Academy of Sciences May 2021. DOI: 10.1073/pnas.2021426118

- Srikar Krishna, Srikala Raghavan*, R. Dasgupta*, Palakodeti D. tRNA-derived fragments (tRFs): establishing their turf in post-transcriptional gene regulation. Cellular and Molecular Life Sciences January 2021. DOI: 10.1007/s00018-020-03720-7

- M. Haroon, Vairavan Lakshmanan, Souradeep R. Sarkar, Kai Lei, P. Vemula*, Palakodeti D. Mitochondrial state determines functionally divergent stem cell population in planaria. Stem cell Reports May 2021. DOI: 10.1016/j.stemcr.2021.03.022

- Alok Javali, Vairavan Lakshmanan, Palakodeti D , R. Sambasivan. Modulation of β‐catenin levels regulates cranial neural crest patterning and dispersal into first pharyngeal arch. Developmental Dynamics May 2020. DOI: 10.1002/dvdy.208

- A. Sarkar, Namita Mukundan, S. Sowndarya, Vinay Dubey, R. Babu, Vairavan Lakshmanan, Kannan Rangiah, M. Panicker, Palakodeti D , S. P. Subramanian, R. Subramanian. Serotonin is essential for eye regeneration in planaria Schmidtea mediterranea. FEBS November 2019. DOI 10.1002/1873-3468.13607

- S. Ganesan, Hamenth Kumar Palani, Vairavan Lakshmanan, N. Balasundaram, A. A. Alex, S. David, Arvind Venkatraman, A. Korula, B. George, P. Balasubramanian, Palakodeti D , N. Vyas, V. Mathews. Stromal cells downregulate miR-23a-5p to activate protective autophagy in acute myeloid leukemia. Cell Death and September. https://www.nature.com/articles/s41419-019-1964-8

- Srikar Krishna, Daniel GR Yim, Vairavan Lakshmanan, Varsha Tirumalai, Judice LY Koh, Jung Eun Park, Jit Kong Cheong, Joo Leng Low, Michelle JS Lim, Siu Kwan Sze, Padubidri Shivaprasad, Akash Gulyani, Srikala Raghavan*, Palakodeti D , Ramanuj DasGupta*. Dynamic expression of tRNA‐derived small RNAs define cellular states. EMBO Reports June 2019. https://www.embopress.org/doi/10.15252/embr.201947789

- Radhika Arasala Rao, Alhad Ashok Ketkar, Neelam Kedia, Vignesh K Krishnamoorthy, Vairavan Lakshmanan, Pankaj Kumar, Abhishek Mohanty, Shilpa Dilip Kumar, Sufi O Raja, Akash Gulyani, Chandra Prakash Chaturvedi, Marjorie Brand, Palakodeti D , Shravanti Rampalli. KMT1 family methyltransferases regulate heterochromatin–nuclear periphery tethering via histone and non‐histone protein methylation. EMBO Reports March 2019. https://www.embopress.org/doi/10.15252/embr.201643260

- Michelle Ninochka D’Souza, Naveen Kumar Chandappa Gowda, Vishal Tiwari, Rosana Ottakandathil Babu, Praveen Anand, Sudhriti Ghosh Dastidar, Randhir Singh, Owen G. James, Bhuvaneish Selvaraj, Rakhi Pal, Arati Ramesh, Sumantra Chattarji, Siddharthan Chandran, Akash Gulyani, Palakodeti D and Ravi S. Muddashetty. FMRP Interacts with C/D Box snoRNA in the Nucleus and Regulates Ribosomal RNA Methylation. iScience 2018 November. https://www.cell.com/iscience/pdf/S2589-0042(18)30197-4.pdf

- Krishna S, Palakodeti D ,Solana J Post-transcriptional regulation in planarian stem cells Seminars in Cell & Developmental Biology 2018 8 June. https://doi.org/10.1016/j.semcdb.2018.05.013

- Peruvemba SS, Babu P, Palakodeti D ,Subramanian R Identification of multiple isomeric core chitobiose-modified high mannose and paucimannose N-glycans in the planarian Schmidtea mediterranea J Biol Chem. 2018 Feb 23. pii: jbc.RA117.000782. doi: 10.1074/jbc.RA117.000782.

- Sasidharan V, Marepally S, Elliott A S, Baid S, Lakshmanan V, Nayyar N, Bansal D, Alvarado AS, Vemula P, Palakodeti D The miR-124 family of microRNAs is crucial for regeneration of the brain and visual system in the planarian Schmidtea mediterranea Development 2017.doi: 10.1242/dev.144758

- Bansal D, Kulkarni J, Nadahalli K, Lakshmanan V, Krishna S, Sasidharan V, Geo J, DilipKumar S, Pasricha R, Gulyani A, Raghavan S, Palakodeti D Cytoplasmic poly (A) binding protein (PABPC2) critically regulates epidermal maintenance and turnover in planarian Schmidtea mediterranea Development 2017. doi: 10.1242/dev.152942

- Boya R, Yadavalli AD, Nikhat S, Kurukuti S, Palakodeti D,Pongubala JM Developmentally regulated higher-order chromatin interactions orchestrate B cell fate commitment Nucleic Acids Research, 17 August 2017. https://doi.org/10.1093/nar/gkx722

- Yim D, Krishna S, Lakshmanan V, Koh J, Park EJ, Cheong KJ, Low LJ, Lim M, Ip J, Nah MJ, Tan I, Iyer G, Guo H, Sze KS, Raghavan S, Palakodeti D DasGupta R Dynamic expression of tRNA-derived small RNAs define cellular states bioRxiv July 1 2017. doi: https://doi.org/10.1101/158501

- Shettigar N, Joshi A,Dalmeida R, Gopalkrishna R, Chakravarthy A, Patnaik S, Mathew M, Palakodeti D, Gulyani A Hierarchies in light sensing and dynamic interactions between ocular and extraocular sensory networks in a flatworm Science Advances vol. 3 no. 7 28 July 2017.

- Arya D, Sachithanandan SP, Ross C, Palakodeti D, Li S, Krishna S.miRNA-182 regulates percentage of myeloid and erythroid cells in chronic myeloid leukemia. Cell Death and Disease. 2017 Jan 12;8(1):e2547.

- Lakshmanan V, Bansal D, Kulkarni J, Poduval D, Krishna S, Sasidharan V, Anand P, Seshasayee A, Palakodeti D. Genome-Wide Analysis of Polyadenylation Events in Schmidtea mediterranea. G3 (Bethesda). 2016 Aug 9. pii: g3.116.031120. doi:10.1534/g3.116.031120. [Epub ahead of print] PubMed PMID: 27489207.

- Natarajan N, Ramakrishnan P, Lakshmanan V, Palakodeti D, Rangiah K. A quantitative metabolomics peek into planarian regeneration. Analyst. 2015 May 21;140(10):3445-64. doi: 10.1039/c4an02037e. Epub 2015 Mar 27. PubMed PMID: 25815385.
- Rao RA, Dhele N, Cheemadan S, Ketkar A, Jayandharan GR, Palakodeti D, Rampalli S. Ezh2 mediated H3K27me3 activity facilitates somatic transition during human pluripotent reprogramming.Sci Rep. 2015 Feb 4;5:8229. doi: 10.1038/srep08229. PubMed PMID: 25648270; PubMed Central PMCID: PMC4316165.

- Vyas N, Walvekar A, Tate D, Lakshmanan V, Bansal D, Lo Cicero A, Raposo G, Palakodeti D, Dhawan J. Vertebrate Hedgehog is secreted on two types of extracellular vesicles with different signaling properties. Sci Rep. 2014 Dec 8;4:7357. doi: 10.1038/srep07357. PubMed PMID: 25483805; PubMed Central PMCID: PMC4258658.

- Rangiah K, Palakodeti D. Comprehensive analysis of neurotransmitters from regenerating planarian extract using an ultrahigh-performance liquid chromatography/mass spectrometry/selected reaction monitoring method. Rapid Commun Mass Spectrom. 2013 Nov 15;27(21):2439-52. doi: 10.1002/rcm.6706. PubMed PMID: 24097401.
- Sasidharan V, Lu YC, Bansal D, Dasari P, Poduval D, Seshasayee A, Resch AM, Graveley BR, Palakodeti D. Identification of neoblast and regeneration-specific miRNAs in the planarian Schmidtea mediterranea. RNA. 2013 Oct;19(10):1394-404. doi: 10.1261/rna.038653.113. Epub 2013 Aug 23. PubMed PMID: 23974438; PubMed Central PMCID: PMC3854530.

- Krishna S, Nair A, Cheedipudi S, Poduval D, Dhawan J, Palakodeti D, Ghanekar Y. Deep sequencing reveals unique small RNA repertoire that is regulated during head regeneration in Hydra magnipapillata. Nucleic Acids Res. 2013 Jan 7;41(1):599-616. doi: 10.1093/nar/gks1020. Epub 2012 Nov 19. PubMed PMID: 23166307; PubMed Central PMCID: PMC3592418.

- Resch AM, Palakodeti D, Lu YC, Horowitz M, Graveley BR. Transcriptome analysis reveals strain-specific and conserved stemness genes in Schmidtea mediterranea. PLoS One. 2012;7(4):e34447. doi: 10.1371/journal.pone.0034447. Epub 2012 Apr 4. PubMed PMID: 22496805; PubMed Central PMCID: PMC3319590.

- Resch AM, Palakodeti D. Small RNA pathways in Schmidtea mediterranea. Int J Dev Biol. 2012;56(1-3):67-74. doi: 10.1387/ijdb.113436ar. Review. PubMed PMID: 22450996.

- Hollenbach JP, Resch AM, Palakodeti D, Graveley BR, Heinen CD. Loss of DNA mismatch repair imparts a selective advantage in planarian adult stem cells. PLoS One. 2011;6(7):e21808. doi: 10.1371/journal.pone.0021808. Epub 2011 Jul 1. PubMed PMID: 21747960; PubMed Central PMCID: PMC3128615.

- Eipper-Mains JE, Kiraly DD, Palakodeti D, Mains RE, Eipper BA, Graveley BR. microRNA-Seq reveals cocaine-regulated expression of striatal microRNAs. RNA. 2011 Aug;17(8):1529-43. doi: 10.1261/rna.2775511. Epub 2011 Jun 27. PubMed PMID: 21708909; PubMed Central PMCID: PMC3153976.

- Lu YC, Smielewska M, Palakodeti D, Lovci MT, Aigner S, Yeo GW, Graveley BR. Deep sequencing identifies new and regulated microRNAs in Schmidtea mediterranea.RNA. 2009 Aug;15(8):1483-91. doi: 10.1261/rna.1702009. Epub 2009 Jun 24. PubMed PMID: 19553344; PubMed Central PMCID: PMC2714757.

- Palakodeti D, Smielewska M, Lu YC, Yeo GW, Graveley BR. The PIWI proteins SMEDWI-2 and SMEDWI-3 are required for stem cell function and piRNA expression in planarians. RNA. 2008 Jun;14(6):1174-86. doi: 10.1261/rna.1085008. Epub 2008 May 2. PubMed PMID: 18456843; PubMed Central PMCID: PMC2390803.

- Palakodeti D, Smielewska M, Graveley BR. MicroRNAs from the Planarian Schmidtea mediterranea: a model system for stem cell biology. RNA. 2006 Sep;12(9):1640-9. Epub 2006 Jul 18. PubMed PMID: 16849698; PubMed Central PMCID: PMC1557691.

Collaborators














Lab Photos







Funding
Our work would not be possible without the generous funding from those listed below
Get in Touch
Join the team!!
JRF/SRF and post-doc applications are welcome !
Interested candidates write to Dr.Dasaradhi Palakodeti
If you are interested in our work and want to contact us.
Contact Details
- Dasaradhi Palakodeti (Work)(Personal)
- +91 80 2366 6001
-
inStem-NCBS, Second Floor, Southern Lab Complex,
GKVK Post, Bellary Road,
Bangalore 560065, India.